Quantitative proteomics is no longer a niche technical choice — it’s a strategic decision that shapes experimental design, budget, statistical power, and ultimately the biological conclusions a laboratory can defend. Choosing between label-free quantitation (LFQ) and labeled approaches (isobaric tags such as TMT and iTRAQ, or metabolic labeling like SILAC) determines where quantitative information is embedded in the workflow, which sources of variability are controlled, and which downstream bioinformatics steps become critical. In this feature I summarize the conceptual landscape, verify core technical claims, highlight strengths and risks, and provide a practical decision matrix that labs can use to match quantitation strategy to scientific goals.
Proteomic workflows have three broad stages relevant to quantitation: sample preparation and labeling (if any), mass spectrometric data acquisition, and computational analysis. The four commonly discussed quantitation strategies—LFQ, TMT / iTRAQ (isobaric tagging), and SILAC (metabolic labeling)—differ precisely in when and how quantitation information is created.
Equip experimental plans with realistic pilot experiments, clear statistical thresholds, and explicit acquisition/analysis SOPs. When those elements are in place, the chosen quantitation strategy will serve the biology — not the other way around — and yield proteomic measurements that are robust, reproducible, and defensible.
Source: Technology Networks Label-Free vs Labeled Proteomics Quantitation Techniques
 Background / Overview
Background / Overview
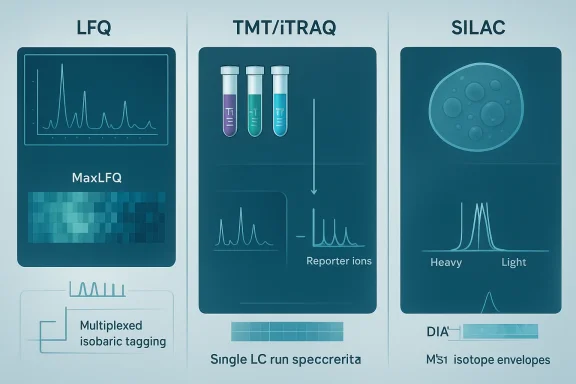
Proteomic workflows have three broad stages relevant to quantitation: sample preparation and labeling (if any), mass spectrometric data acquisition, and computational analysis. The four commonly discussed quantitation strategies—LFQ, TMT / iTRAQ (isobaric tagging), and SILAC (metabolic labeling)—differ precisely in when and how quantitation information is created.- LFQ compares peptide signal intensities across independent LC–MS/MS runs and relies heavily on run alignment, normalization, and missing-value handling during data analysis.
- Isobaric tagging (TMT/iTRAQ) chemically labels peptides from separate samples, pools them, and derives relative quantitation from reporter ions observed during MS/MS, reducing run-to-run variability and enabling multiplexing.
- SILAC introduces stable isotope–labeled amino acids during cell growth so that labeled and unlabeled peptide pairs are mixed early and quantified at the MS1 level.
Label‑Free Quantitation (LFQ): flexibility at the price of computation
How LFQ works
LFQ typically measures peptide abundance via integrated MS1 signal intensity (extracted ion chromatograms) or, in simpler implementations, spectral counts. Modern LFQ pipelines align peptide features across runs and normalize intensities so that a peptide’s integrated peak area becomes a proxy for its relative abundance across samples. Advanced LFQ algorithms (for example, MaxLFQ) reconstruct protein-level abundance profiles by maximizing the use of pairwise peptide ratios across runs and applying delayed normalization to reduce bias. This approach scales naturally to very large cohorts because samples are not pooled prior to acquisition.Strengths
- Scalability and flexibility: There is no intrinsic upper limit to the number of independent samples LFQ can compare, so it suits large clinical cohorts, longitudinal studies, and exploratory experiments.
- Lower reagent costs: No chemical tags or metabolic labels required — consumable costs are lower than labeled workflows.
- Compatibility: Works with tissues, biofluids, and any sample that cannot be metabolically labeled.
Weaknesses and risks
- Run-to-run variability: Because every sample is analyzed in a separate LC–MS/MS injection, instrument drift, chromatography differences, and subtle changes in ionization can confound comparisons unless carefully corrected.
- Missing values are common: Missing peptide quantifications (non-detects) are a persistent challenge in LFQ datasets, sometimes affecting substantial fractions of the data; appropriate imputation or modeling is therefore essential to avoid bias. Comparative studies of imputation methods show that missing values in LFQ are complex (MAR and MNAR) and that methods such as random-forest (RF) or local least squares (LLS) imputation often outperform simpler strategies.
- Lower precision for small fold changes: Detecting small expression changes across many samples can be harder in LFQ than in labeled within-run comparisons.
Practical notes
- Use established pipelines (MaxQuant/MaxLFQ + Perseus or comparable tools) for large LFQ studies; MaxLFQ was explicitly designed to scale to hundreds of samples and to maximize precision by using maximal peptide ratio extraction.
- Plan for robust QC: randomized run order, repeated technical injections, and internal standards (e.g., peptide or protein spike-ins) substantially help downstream normalization.
Isobaric labeling (TMT and iTRAQ): multiplexed precision with caveats
How isobaric tagging works
Isobaric tags chemically label peptides from different samples so that all labeled peptides are isobaric (same intact mass) and co-elute during LC. During MS/MS fragmentation, each tag releases a reporter ion of a unique mass; relative reporter intensities quantify sample contributions within the pooled run. The approach embeds quantitation within each MS/MS event, so all samples in the multiplex experience the same chromatography and ionization conditions.Strengths
- Multiplexing and instrument efficiency: Multiple samples (commonly 6–18, depending on chemistry and instrument) are compared within a single LC–MS/MS run, which reduces run-to-run variance and improves throughput. Newer TMTpro reagents support up to 18 channels (18-plex), enabling high within-run comparisons.
- Fewer missing values: Because peptides from all channels are present in the same MS run, missing reporter values are much less common than missing identifications across separate LFQ runs.
- Improved precision for small fold changes: Analyzing samples together reduces many technical sources of variation, improving the detection of subtle abundance differences.
Key technical risk: ratio compression
When co-isolated contaminant peptides (from other precursors within the isolation window) are fragmented along with the target peptide, reporter ion intensities become mixtures of signals. This “co-isolation interference” leads to ratio compression, underestimating real fold changes. The problem is well documented and can meaningfully bias results if unmitigated. Strategies to mitigate ratio compression include:- Narrow isolation windows and optimized acquisition parameters.
- MS3-based methods (SPS-MS3) that perform an extra round of isolation and fragmentation so reporter ions are measured from purer fragment populations; this approach has been shown to largely eliminate ratio distortion in controlled tests.
- Computational correction and post-acquisition algorithms to model and correct interference.
Practical considerations
- If you need within-run multiplexing (e.g., controlled cohorts, time-course replicates, or single-cell multiplexing strategies), TMT/iTRAQ provide major advantages in precision and instrument economy.
- Budget accordingly: isobaric reagents are expensive and add complexity to sample prep.
- For experiments where ratio compression would catastrophically affect conclusions (large background contamination or very complex samples), plan to use MS3 or advanced acquisition strategies and validate with spike-ins or orthogonal methods.
SILAC (metabolic labeling): accurate early combination, limited scope
How SILAC works
SILAC labels proteins by culturing cells in media containing heavy or light isotopic amino acids. Labeled and unlabeled cell populations are combined early — often before lysis — and processed together. Quantitation derives from paired isotope envelopes at the MS1 level; because the pairs co-elute and are measured together, many technical variabilities are removed.Strengths
- Very high quantitative accuracy: Early mixing neutralizes many sources of technical bias because heavy/light peptide pairs experience identical processing and acquisition.
- Simple chemistry: Label incorporation happens through growth media rather than a chemical derivatization step, often making workflows experimentally convenient.
Limitations
- Applicable only to living cultures: SILAC requires metabolic incorporation and thus is restricted to cell culture systems (or specially labeled small organisms) — tissues, primary clinical samples, and most biofluids cannot be analyzed by SILAC.
- Limited multiplexing: Traditional SILAC experiments are commonly duplex or triplex; increasing channels is nontrivial.
- Label incorporation challenges: Slow-growing cells or systems that synthesize amino acids de novo may require extended labeling time or yield incomplete incorporation.
- MS1-based quantitation reduces identification space: Because quantitation is MS1-based, achieving deep proteome coverage can be constrained relative to some other methods.
Practical notes
- Use SILAC when cell-culture system biology is the focus and when absolute accuracy for relative changes is paramount.
- Confirm incorporation completeness with pilot experiments, and account for cost/time of multiple cell passages.
Where the quantitation happens: MS1 vs MS2 and why it matters
- LFQ: quantitation at MS1 across runs; depends on peptide feature alignment, XIC area integration, and robust normalization. MaxLFQ is a prominent algorithmic solution that maximizes peptide ratio usage to produce precise protein-level quantitation across large datasets.
- Isobaric tags (TMT/iTRAQ): quantitation at MS2 (reporter ions) within runs; samples pooled and compared simultaneously, improving precision but introducing co-isolation interference risk (ratio compression) that can be mitigated with MS3 or other acquisition strategies.
- SILAC: quantitation at MS1 within the same run using isotope pairs, offering high accuracy but limited multiplex capacity and sample-type compatibility.
Alternatives and the growing role of DIA/SWATH
A mature third axis of choice exists outside the classic LFQ vs labeled dichotomy: data-independent acquisition (DIA), including SWATH-MS. DIA collects fragment-ion data for wide precursor windows, creating a permanent digital map of fragment spectra that can later be queried with spectral libraries or library‑free algorithms. DIA combines many benefits — improved reproducibility, fewer missing values, and retrospective re‑querying of data — and it’s increasingly used in both label-free and targeted clinical applications. For projects that need consistent quantitation across many samples with lower missingness than classic DDA/LFQ, DIA is a compelling alternative.Comparative strengths and a decision matrix
Below is a compact decision matrix to help match strategy to experiment type.- If you have large sample numbers, diverse sample types (tissues, biofluids), or a limited reagent budget: favor LFQ (with careful QC and robust imputation strategies).
- If you need high precision across a defined set of conditions, want to minimize missing values, and can afford reagents and instrument time: favor isobaric tagging (TMT/iTRAQ); plan for interference mitigation (MS3 or narrow isolation) if small fold changes are critical.
- If your system is cell-culture compatible and you prioritize early sample combination and maximum quantitative accuracy: consider SILAC, acknowledging multiplex and sample constraints.
- If you want reproducible, archival-quality runs with fewer missing values and the ability to re‑mine data later: evaluate DIA / SWATH as a modern label-free alternative.
- Define the minimum effect size you must detect and the acceptable false-discovery risk.
- List sample types and whether metabolic labeling is feasible.
- Quantify available instrument time and access (e.g., many short runs vs fewer multiplexed runs).
- Budget for reagents plus data storage and analysis infrastructure.
- Plan analytical pipelines (MaxQuant, Proteome Discoverer, DIA‑NN, Spectronaut, Perseus) and compute resources for imputation/normalization.
- Design pilots that include internal standards and spike‑ins to validate chosen approach.
Best practices, QC and data analysis recommendations
- Randomize sample order and block by batch. For LFQ, distribute biological groups evenly across LC batches to avoid confounding.
- Include technical replicates and pooled reference samples. For isobaric designs, include a common reference channel when sample numbers exceed plex size.
- For TMT/iTRAQ, adopt MS3/SPS-MS3 or optimized narrow‑isolation acquisition strategies when ratio compression threatens conclusions; validate using mixture experiments (two-proteome designs) or spike‑ins. MS3 has been shown to markedly reduce reporter distortion in controlled tests.
- For LFQ, pick a robust feature alignment and normalization tool (MaxLFQ is widely validated) and plan imputation strategy up front; comparative analyses show RF and LLS imputation often perform best for LFQ missing data.
- For DIA, invest in spectral library generation or use high-quality library‑free tools; DIA workflows can produce much higher completeness and reproducibility for large cohorts.
- Report technical parameters transparently: isolation width, MS1/ MS2 resolution, AGC targets, injection times, and any MS3 settings — these acquisition choices materially affect quantitative accuracy and reproducibility. Several cross-platform studies emphasize the need to optimize instrument parameters to minimize interference without sacrificing identifications.
Cost and throughput: an operational reality check
Be transparent with stakeholders: isobaric reagents are a nontrivial per-sample cost and require additional hands-on time. LFQ minimizes reagent spend but may require more instrument time and more sophisticated downstream computational resources to handle missingness and batch corrections. SILAC implies additional biological costs (label media and extended cell passages) and is limited to lab-amenable systems. Exact reagent prices and instrument rates vary widely across regions and procurement channels; treat any single price estimate as approximate and re‑quote at procurement. Where possible, build cost comparisons that include reagent, instrument, and analyst time rather than reagent costs alone.Emerging directions and caveats
- Hyperplexing strategies and novel isobaric chemistries continue to expand channel capacity (for example TMTpro reagents reaching 16–18 channels), but higher plex does not automatically mean better results — interference, dynamic range, and instrument duty cycle still shape outcome quality.
- Single‑cell proteomics increasingly leverages isobaric multiplexing plus carrier channels and real-time search MS3 strategies to boost depth at the single-cell level; these specialized methods require careful benchmarking because carrier-channel artifacts and ratio-compression effects differ from bulk experiments.
- Computational advances (improved imputation, ratio‑compression correction algorithms, and library‑free DIA analysis) are changing the landscape. Still, laboratory validation (spike-ins, orthogonal assays) remains the gold standard for asserting quantitative claims.
Final recommendations — a decision workflow
- Step 1: Clarify the biological question and minimum detectable fold change.
- Step 2: Assess sample type compatibility (can you use SILAC? must you analyze tissues?).
- Step 3: Estimate cohort size and instrument access (many single runs vs few multiplexed runs).
- Step 4: Run a small technical pilot: include pooled references, spike‑ins at expected effect sizes, and test both acquisition and analysis pipelines.
- Step 5: Choose the approach that minimizes confounding while delivering required precision; validate with orthogonal measurements as available.
Conclusion
There is no universally “best” quantitative proteomics method; each approach embeds quantitation in a different place in the workflow and trades off flexibility, precision, cost, and sample compatibility. LFQ wins on scalability and low reagent cost but demands rigorous computational handling of variability and missing values. Isobaric labeling (TMT/iTRAQ) wins on multiplexed precision and instrument efficiency but requires careful attention to ratio compression and often higher consumable expense; MS3/SPS-MS3 and optimized acquisition mitigate interference but change duty cycles. SILAC offers best-in-class early-sample-mixing accuracy but is confined to cell-culture systems. Meanwhile, DIA/SWATH is emerging as a powerful label-free option that combines reproducibility and completeness, and deserves consideration where archival digital maps and low missingness matter.Equip experimental plans with realistic pilot experiments, clear statistical thresholds, and explicit acquisition/analysis SOPs. When those elements are in place, the chosen quantitation strategy will serve the biology — not the other way around — and yield proteomic measurements that are robust, reproducible, and defensible.
Source: Technology Networks Label-Free vs Labeled Proteomics Quantitation Techniques